BLAST Tool
The BLAST Tool
The Basic Local Alignment Search Tool (BLAST) is a software tool for comparing biological sequences (either nucleotides or proteins) against sequences databanks in order to locate homologs between sequences. Since the BLAST tool is very useful for scientists to identify the biological roles of new sequences, its has been integrated in LabCollector LIMS.
Run a NCBI Blast or an internal Blast on LabCollector LIMS!
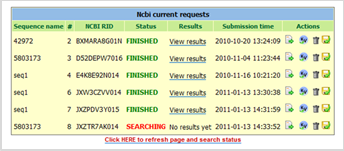
BLAST search jobs can now be directly run from within the Sequence Add-on of LabCollector. This new service enables you to run search jobs using your own sequences as queries and/or target databanks. Two search services are available: NCBI Blast Server or LabCollector Local Blast. 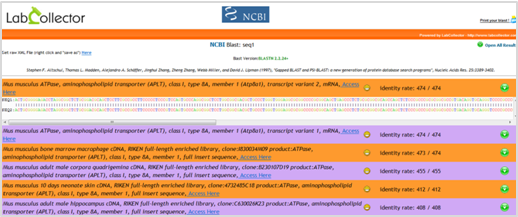 Whichever the search services used, LabCollector LIMS automatically handles the whole search processes. Results are stored within the LabCollector database and can be viewed either graphically or in plain text.
Whichever the search services used, LabCollector LIMS automatically handles the whole search processes. Results are stored within the LabCollector database and can be viewed either graphically or in plain text.